Variant TP53BP1 rs560191 G>C is associated with risk of gastric cardia adenocarcinoma in a Chinese Han population
Introduction
Gastric cardia adenocarcinoma (GCA) is one of the most common malignant tumors and is among the leading causes of cancer-related death. GCA has increased dramatically in North America and Western European countries (1). In China, GCA shares very similar geographic distribution with esophageal squamous cell carcinoma (ESCC) epidemiologically (1,2). Both genetic and environmental factors may contribute to the etiology of GCA (3). Single nucleotide polymorphisms (SNPs), accounting for >90% of genetic variations, may influence the function of genes (4). SNPs have been suggested to affect individual differences in disease susceptibility (5).
Apoptosis is a crucial mechanism against hyper-proliferation and malignancy, and has been considered as a fundamental component in cancer pathogenesis (6). Caspase8 (CASP8), Caspase7 (CASP7), tumor protein 53-binding protein 1 (TP53BP1) and C1orf10/Cornulin (CRNN) are among key genes related to apoptosis. CASP8 is an initiator CASP and a key regulator of apoptosis, which plays an important role in cancer development and progression (7). In critical cell structures, CASP7 conducts a coordinated program of proteolysis and resulting in destruction, acting as an executioner CASP (8). TP53BP1 interacts specifically with p53 and helps mediate the DNA damage checkpoint (9). TP53BP1 participates in both DNA repair and cell cycle control. TP53BP1 contains two BRCA1 C-terminal (BRCT) domains, which are essential for its tumor suppressor functions (10). Available functional data also suggests that the C1orf10/CRNN protein might protect against apoptosis during cellular responses to stress (11).
In a previous study, we investigated the associations between functional polymorphisms TP53BP1 rs560191 G>C, CASP8 rs1035142 G>T, CASP7 rs3127075 G>C, CASP7 rs7907519 C>A and six C1orf10/CRNN variants and ESCC susceptibility in a hospital-based case-control study (12). In this study, we hypothesize that these SNPs may alter individual risk of GCA. Thus, we performed genotyping analyses for the 10 SNPs using 243 GCA cases and 476 controls in a Chinese Han population.
Patients and methods
Ethical approval of study protocol
The Review Board of Jiangsu University (Zhenjiang, China) approved this study (Zhenjiang, China). All subjects included in the study gave written informed consent. We have complied with the World Medical Association Declaration of Helsinki regarding ethical conduct of research involving human subjects and/or animals.
Study subjects
From October 2008 to July 2010, 243 GCA patients contained in the case group were consecutively recruited at the Affiliated People’s Hospital of Jiangsu University and Affiliated Hospital of Jiangsu University (Zhenjiang, China). For all the cases, pathological examination was used to make a definite diagnosis. Tumor, nodal, and metastatic (TNM) stage was according to the seventh edition of the Union for International Cancer Control (UICC) TNM staging (13). The exclusion criteria included: patients who previously had cancer or any metastatic cancer, received radiotherapy or chemotherapy. A total of 476 cancer-free controls were contained in this study, and among which 380 controls were recruited from the two hospitals aforementioned during the same time period. Another 96 controls were recruited from hospitals in Changzhou city (which is adjacent to Zhenjiang) as previous described (14). The controls were matched to the cases in terms of age (±5 years) and sex. Most of the controls were admitted to the hospitals to receive treatment for trauma.
Using a pre-tested questionnaire, each patient was personally questioned by trained investigator to obtain information on demographic data (e.g., age, sex) and related environmental risk factors (including tobacco using and alcohol consumption). Individuals who smoked one cigarette per day for >1 year were considered to be “smokers”. Subjects who consumed ≥3 alcoholic drinks a week for >6 months were defined as “alcohol drinkers”. After the investigation, 2 mL venous blood sample was collected from each subject.
DNA extraction and genotyping analysis
Blood samples were collected from subjects using Vacutainers and drawn into tubes lined with ethylenediamine tetra-acetic acid (EDTA). The QIAamp DNA Blood Mini Kit (Qiagen, Berlin, Germany) was used to extract genomic DNA from peripheral blood (15). Sample DNA (10 ng) was amplified by PCR according to the manufacturer’s recommendations. A custom-by-design 48-Plex SNPscanTM Kit (Genesky Biotechnologies Inc., Shanghai, China) was used to determine genotypes of the ten SNPs as previously described (1,16-18). This kit was developed according to patented SNP genotyping technology by Genesky Biotechnologies Inc., which was substantially based on double ligation and multiplex fluorescence PCR. Repeated analyses were accomplished to guarantee the genotyping quality by randomly choosing 4% of samples with high DNA quality.
Statistical analysis
Differences in the distributions of demographic characteristics, selected variables, and genotypes of the TP53BP1 rs560191 G>C, CASP8 rs1035142 G>T, CASP7 rs3127075 G>C, CASP7 rs7907519 C>A and six C1orf10/CRNN variants between the cases and controls were estimated using the χ2 test. The associations between the ten SNPs and susceptibility of GCA were evaluated by calculating the odds ratios (ORs) and their 95% confidential intervals (95% CIs) using logistic regression analyses for crude ORs and adjusted ORs when adjusting for age, sex, smoking and drinking status. The Hardy-Weinberg equilibrium (HWE) was tested by a goodness-of-fit χ2 test to compare the observed genotype frequencies to the expected ones among the control group. All statistical analyses were performed with SAS 9.1.3 (SAS Institute, Cary, NC, USA). P<0.05 was considered statistically significant.
Results
Characteristics of study population
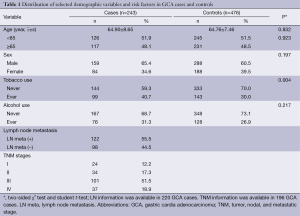
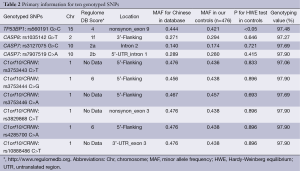
Characteristics of cases and controls included in the study are summarized in Table 1. The cases and controls appeared to be adequately matched on age and sex as suggested by the χ2 tests (P=0.923 and P=0.197, respectively). As shown in Table 1, no significant difference was detected on drinking status between the cases and the controls (P=0.217), but smoking rate was higher in GCA patients than in control subjects (P=0.004). Lymph node metastasis information was available in 220 (90.5%) of 243 GCA patients; regional lymph node metastasis was present in 122 (55.5%) cases. TNM stage data were available in 196 (80.7%) of 243 patients (stage I: 24; stage II: 34; stage III: 101, stage IV: 37). The primary information for ten genotyped SNPs is shown in Table 2. For the ten SNPs, the genotyping was successful ranging from 97.06% to 97.90% in all 719 samples. The concordance rates of repeated analyses were 100%. Minor allele frequency (MAF) in our controls was similar to MAF for Chinese in database for all ten SNPs. The observed genotype frequencies for these ten polymorphisms in the controls were in HWE except TP53BP1 rs560191 G>C (Table 2).
Full table
Full table
Associations between ten polymorphisms and risk of GCA
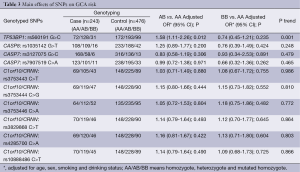
The genotype distributions of TP53BP1 rs560191 G>C, CASP8 rs1035142 G>T, CASP7 rs3127075 G>C, CASP7 rs7907519 C>A and six C1orf10/CRNN in the cases and the controls are shown in Table 3. In the single locus analyses, the genotype frequencies of TP53BP1 rs560191 G>C were 31.17% (GG), 55.41% (GC), 13.42% (CC) in the case patients and 37.07% (GG), 41.59% (GC), 21.34% (CC) in the control subjects, and the difference was statistically significant (P=0.001). When the TP53BP1 rs560191 GG homozygote genotype was used as the reference group, the GC genotype was associated with a significantly increased risk for GCA (GC vs. GG: adjusted OR =1.58, 95% CI =1.11-2.26, P=0.012). The CC genotype was not associated with the risk of GCA compared with the GG genotype (CC vs. GG: adjusted OR =0.74, 95% CI =0.45-1.21, P=0.235) (Table 3).
Full table
None of the CASP8 rs1035142 G>T, CASP7 rs3127075 G>C, CASP7 rs7907519 C>A and six C1orf10/CRNN polymorphisms achieved a significant difference in the genotype distributions between the cases and the controls. Logistic regression analyses revealed that these nine polymorphisms were not associated with the risk of GCA (Table 3). Bonferroni correction was performed to correct the P value; Pcorrect>0.05 in all comparison models for the ten polymorphisms (Table 3).
Stratification analyses of TP53BP1 rs560191 G>C polymorphism and risk of GCA
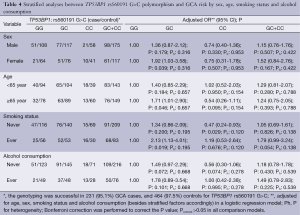
To evaluate the effects of TP53BP1 rs560191 G>C genotypes on GCA risk according to different age, sex, smoking and alcohol drinking status; we performed the stratification analyses. A significantly increased risk of GCA associated with the TP53BP1 rs560191 G>C polymorphism was evident among female patients, older patients and patients who smoked. A significantly decreased risk of GCA associated with the TP53BP1 rs560191 G>C polymorphism was evident among patients who never smoked (Table 4). Bonferroni correction was performed to correct the P value; Pcorrect>0.05 in all comparison models (Table 4).
Full table
Discussion
We conducted this study to investigate the associations of TP53BP1 rs560191 G>C, CASP8 rs1035142 G>T, CASP7 rs3127075 G>C, CASP7 rs7907519 C>A and six C1orf10/CRNN SNPs with susceptibility to GCA in a high risk Chinese population. Our findings revealed that the variant TP53BP1 rs560191 GC genotype was associated with a significantly increased risk of GCA.
TP53BP1 interacts specifically with p53, which was a well known tumor-associated gene (19). TP53BP1 binds to p53 and plays a role in responses to DNA damage. A study in Japan revealed that TP53BP1 rs560191 G>C and p53 Arg72Pro polymorphism had significant interaction in lung cancer risk (20). By cooperating with damage sensors and signal transducers, TP53BP1 helps mediate the DNA damage checkpoint (9). TP53BP1 plays an important role in both DNA repair and cell cycle control (21). TP53BP1 contains two BRCT domains, which are essential for tumor suppressor functions (10). TP53BP1 SNPs may play an important role in the etiology of cancer because of the direct role of TP53BP1 in the cellular response to DNA damage. Previous studies revealed no association between TP53BP1 rs560191 G>C (Asp353Glu) SNPs and cancer risk (22-27); however, Kiyohara reported that the TP53BP1 rs560191 G>C SNP was associated with a decreased risk of lung cancer (20). In previous a meta-analysis, no significant association between TP53BP1 rs560191 G>C SNP and cancer risk was found in the overall population (28).
The negative results for an association of TP53BP1 rs560191 G>C SNP and cancer risk in these previous case-control reports and meta-analyses are not in accordance with our results. However, our stratified analysis revealed that the increased risk was largely attributed to female patients, older patients and patients who smoked. Among patients who never smoked, the SNP was protective, indicating age, sex and environment differences might play an important role in cancer etiology related to the TP53BP1 rs560191 G>C SNP. Previously, it was found that neither TP53BP1 rs560191 G>C polymorphism nor CASP7 rs3127075 G>C, CASP7 rs7907519 C>A and six C1orf10/CRNN are associated with the risk of ESCC, which is in accordance with our results.
The frequencies of genetic polymorphisms often vary between ethnic groups. In the current study involving Chinese subjects, the allele frequency of TP53BP1 rs560191 C was 0.421 among 476 control subjects, which is consistent with the Chinese Han population (0.444), but lower than Sub-Saharan African population (1.000) and higher than the European population (0.308), according to the SNP DataBase (http://hapmap.ncbi.nlm.nih.gov/).
Using Power and Sample Size Calculation (PS, version 3.0, 2009, http://biostat.mc.vanderbilt.edu/twiki/bin/view/Main/PowerSampleSize), and considering TP53BP1 rs560191 G>C mutant alleles in the control group, OR, GCA samples and control sample sizes, the power of our analysis (α=0.05) was 0.809 in the 231 GCA cases and 464 controls with an OR =1.58 for TP53BP1 rs560191 G>C.
This study has some limitations. First, the subjects may not be representative because our study was based on hospital attendees instead of the general population. False positive results may be obtained because of inherited biases. Second, the statistical power of our study was limited because of the moderate sample size and lack of a validation cohort. Some significant findings may have occurred by chance. Third, single SNP locus analyses may not provide a comprehensive understanding of the genetic effects of these candidate genes; further fine-mapping analysis of the susceptibility region is warranted. Combinations of certain genotypes may be also more discriminating as risk factors than single locus genotypes. Finally, because of the lack of detailed information on cancer metastasis and survival, further analyses on the role of TP53BP1 polymorphisms in GCA progression and prognosis cannot be achieved.
Conclusions
We found an association between genetic variant TP53BP1 rs560191 G>C and susceptibility to GCA. However, the sample size is not large enough to draw precise conclusions. Further well-designed studies using a large validation set are needed to support our findings.
Acknowledgements
We wish to thank Dr. Da Ding and Dr. Yan Liu (Genesky Biotechnologies Inc., Shanghai, China) for technical support. This study was supported in part by National Natural Science Foundation of China (No. 81101889).
Disclosure: The authors declare no conflict of interest.
References
- Wang LD, Zheng S, Zheng ZY, et al. Primary adenocarcinomas of lower esophagus, esophagogastric junction and gastric cardia: in special reference to China. World J Gastroenterol 2003;9:1156-64. [PubMed]
- Pera M, Cameron AJ, Trastek VF, et al. Increasing incidence of adenocarcinoma of the esophagus and esophagogastric junction. Gastroenterology 1993;104:510-3. [PubMed]
- Zhang L, Du C, Guo X, et al. Interleukin-8-251A/T polymorphism and Helicobacter pylori infection influence risk for the development of gastric cardiac adenocarcinoma in a high-incidence area of China. Mol Biol Rep 2010;37:3983-9. [PubMed]
- Hu S, Song QB, Yao PF, et al. No relationship between IL-1B gene polymorphism and gastric acid secretion in younger healthy volunteers. World J Gastroenterol 2005;11:6549-53. [PubMed]
- Zhang W, Li C, Wang J, et al. Functional polymorphisms in FAS/FASL system contribute to the risk of occurrence but not progression of gastric cardiac adenocarcinoma. Hepatogastroenterology 2012;59:141-6. [PubMed]
- Hengartner MO. The biochemistry of apoptosis. Nature 2000;407:770-6. [PubMed]
- Krelin Y, Zhang L, Kang TB, et al. Caspase-8 deficiency facilitates cellular transformation in vitro. Cell Death Differ 2008;15:1350-5. [PubMed]
- Lee WK, Kim JS, Kang HG, et al. Polymorphisms in the Caspase7 gene and the risk of lung cancer. Lung Cancer 2009;65:19-24. [PubMed]
- Iwabuchi K, Li B, Massa HF, et al. Stimulation of p53-mediated transcriptional activation by the p53-binding proteins, 53BP1 and 53BP2. J Biol Chem 1998;273:26061-8. [PubMed]
- Williams RS, Green R, Glover JN. Crystal structure of the BRCT repeat region from the breast cancer-associated protein BRCA1. Nat Struct Biol 2001;8:838-42. [PubMed]
- Darragh J, Hunter M, Pohler E, et al. The calcium-binding domain of the stress protein SEP53 is required for survival in response to deoxycholic acid-mediated injury. FEBS J 2006;273:1930-47. [PubMed]
- Yin J, Tang W, Shao A, et al. Caspase8 rs1035142 G>T polymorphism was associated with an increased risk of esophageal cancer in a Chinese population. Mol Biol Rep 2014;41:2037-43. [PubMed]
- Kwon SJ. Evaluation of the 7th UICC TNM Staging System of Gastric Cancer. J Gastric Cancer 2011;11:78-85. [PubMed]
- Cheng J, Zhang H, Zhuang C, et al. Peptidylarginine deiminase type 4 and methyl-CpG binding domain 4 polymorphisms in Chinese patients with rheumatoid arthritis. J Rheumatol 2012;39:1159-65. [PubMed]
- Gu H, Ding G, Zhang W, et al. Replication study of PLCE1 and C20orf54 polymorphism and risk of esophageal cancer in a Chinese population. Mol Biol Rep 2012;39:9105-11. [PubMed]
- Wei J, Zheng L, Liu S, et al. MiR-196a2 rs11614913 T>C polymorphism and risk of esophageal cancer in a Chinese population. Hum Immunol 2013;74:1199-205. [PubMed]
- Li Q, Yin J, Wang X, et al. B-cell Lymphoma 2 rs17757541 C>G polymorphism was associated with an increased risk of gastric cardiac adenocarcinoma in a Chinese population. Asian Pac J Cancer Prev 2013;14:4301-6. [PubMed]
- Sun JM, Li Q, Gu HY, et al. Interleukin 10 rs1800872 T>G polymorphism was associated with an increased risk of esophageal cancer in a Chinese population. Asian Pac J Cancer Prev 2013;14:3443-7. [PubMed]
- Song Y, Li L, Ou Y, et al. Identification of genomic alterations in oesophageal squamous cell cancer. Nature 2014;509:91-5. [PubMed]
- Kiyohara C, Horiuchi T, Miyake Y, et al. Cigarette smoking, TP53 Arg72Pro, TP53BP1 Asp353Glu and the risk of lung cancer in a Japanese population. Oncol Rep 2010;23:1361-8. [PubMed]
- Miwa S, Tome Y, Yano S, et al. Single cell time-lapse imaging of focus formation by the DNA damage-response protein 53BP1 after UVC irradiation of human pancreatic cancer cells. Anticancer Res 2013;33:1373-7. [PubMed]
- Chen K, Hu Z, Wang LE, et al. Polymorphic TP53BP1 and TP53 gene interactions associated with risk of squamous cell carcinoma of the head and neck. Clin Cancer Res 2007;13:4300-5. [PubMed]
- Frank B, Hemminki K, Bermejo JL, et al. TP53-binding protein variants and breast cancer risk: a case-control study. Breast Cancer Res 2005;7:R502-5. [PubMed]
- Zhang H, Hao S, Zhao J, et al. Common genetic variants in 53BP1 associated with nonsmall-cell lung cancer risk in Han Chinese. Arch Med Res 2014;45:84-9. [PubMed]
- Oliveira S, Ribeiro J, Sousa H, et al. Genetic polymorphisms and cervical cancer development: ATM G5557A and p53bp1 C1236G. Oncol Rep 2012;27:1188-92. [PubMed]
- Naidu R, Har YC, Taib NA. Genetic polymorphisms of TP53-binding protein 1 (TP53BP1) gene and association with breast cancer risk. APMIS 2011;119:460-7. [PubMed]
- Ma H, Hu Z, Zhai X, et al. Joint effects of single nucleotide polymorphisms in P53BP1 and p53 on breast cancer risk in a Chinese population. Carcinogenesis 2006;27:766-71. [PubMed]
- Liu L, Jiao J, Wang Y, et al. Lack of association of the TP53BP1 Glu353Asp polymorphism with risk of cancer: a systematic review and meta-analysis. PLoS One 2014;9:e90931. [PubMed]


